Applicant
Prof. Dr. Bernd Meyer
Interdisciplinary Center for Molecular Materials (ICMM)
and Computer Chemistry Center (CCC)
Friedrich-Alexander Universität Erlangen-Nürnberg
Project Summary
We have revised the ultrasoft pseudopotential (USPP) branch of the ab initio molecular dynamics code CPMD. The most important improvements are: (i) A hybrid MPI+OpenMP parallelization has been added to all USPP-specific routines. (ii) Overlapping computation and communication is now used in the most communication-intensive routines. The necessary partitioning of the workload is optimized by auto-tuning algorithms. (iii) The largest global MPI Allreduce operations are replaced by highly tuned node-local parallelized operations using MPI shared-memory windows to avoid inter-node communication. (iv) By introducing a batched algorithm, the throughput of the MPI Alltoall communication and the scalability of the multiple 3d-FFTs are highly improved, which is also beneficial for the norm-conserving pseudopotential (NCPP) code branch of CPMD.
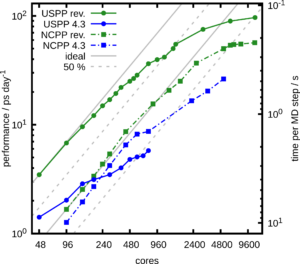
The enhanced performance and scalability of CPMD are demonstrated on SuperMUC-NG at LRZ Garching for the example of a simulation with 256 water molecules. For a single node (48 cores), the revised USPP implementation outperforms the old code by a factor of 2.5. Already at 6 nodes, the parallel efficiency of the old USPP code drops below 50 %, which is often considered a threshold to use the HPC facilities at computing centers. The new code stays above this threshold up to 32 nodes and outperforms the old code by more than an order of magnitude. The achieved performance is about 55 ps of simulation time per day or less than 0.23 s per MD step.
The NCPP calculations also benefit from our new batched FFT. For node counts up to 12, the performance improvement of the NCPP code branch is about 30 % and increases to more than 50 % at 108 nodes due the almost ideal scaling of the new FFT algorithm. While the old USPP implementation was faster than NCPP only up to 4 nodes, it now outperforms NCPP for all node counts by about a factor of 5. This is close to the maximum possible speed-up of 5.7, which is determined by the ratio of the used plane wave basis functions.
Full details on the code improvements and additional benchmarks are given in:
T. Klöffel, G. Mathias, B. Meyer, Integrating state of the art compute, communication, and autotuning strategies to multiply the performance of ab initio molecular dynamics on massively parallel multi-core supercomputers, Comput. Phys. Commun. 260 (2021) 107745; http://dx.doi.org/10.1016/j.cpc.2020.107745

